Hi guys~
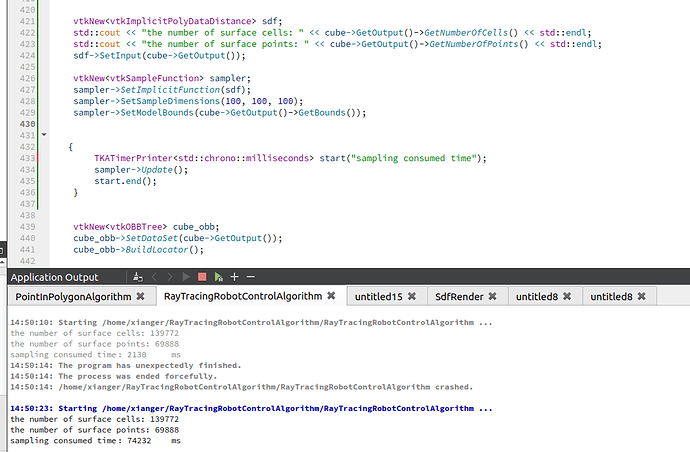
I want to calculate the signed distance field of a polydata, which has about 70000 points and 140000 cells, AND i set the sampling dimension with 100 * 100 * 100. It takes more than 60 seconds, is there existing any another methods to improve the performance? THANK YOU!
CODE BELOW
Hello,
Well, 100x100x100 means a 1M-cell sampling grid. Evaluating the distance to 70k points over 1M cells in 60s is fairly fast for me. One way to improve considerably the performance for that kind of task is to set a search neightborhood (there is no point in including a sample in the far side of the universe to your calculation), but vtkSampleFunction doesn’t seem to have such feature. Another simple way to speed things up is to reduce resolution (say, to 80x80x80).
Check whether vtkSampleFunction uses all your cores during the computation. If it runs single-threaded, one possible way is to distribute some Z-slices to some threads and try to perform the calculation in parallel. For example, run 10 x 100x100x10 vtkSampleFunctions in 10 threads (if your machine has 10 logical processing cores available), then somehow copy the 10 results into a 11th 100x100x100 vtkSampleFunction or other regular-grid-like object (e.g. vtkImageData. vtkStructuredGrid, etc.) after all threads have finished.
regards,
Paulo