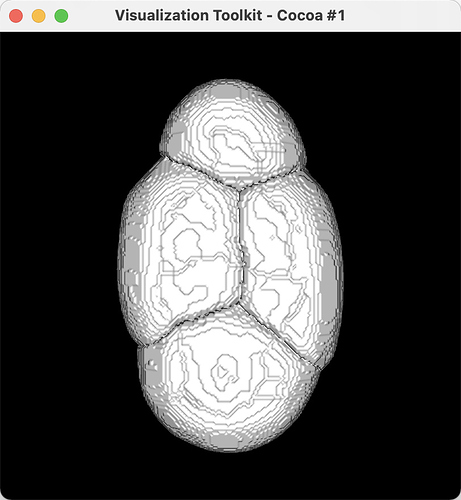
I did a cell segmentation program, and the results of the segmentation were marked. But in 3D reconstruction, I don’t know how to mark the result I marked with different colors and color the target object. 
here is my code for 3D reconstruction and ‘path’ is my image folder.
aRenderer = vtk.vtkRenderer()
renWin = vtk.vtkRenderWindow()
renWin.AddRenderer(aRenderer)
iren = vtk.vtkRenderWindowInteractor()
iren.SetRenderWindow(renWin)
v16 = vtk.vtkNIFTIImageReader()
v16.SetDataByteOrderToLittleEndian()
v16.SetFileName(path)
mc = vtk.vtkMarchingCubes()
mc.SetInputConnection(v16.GetOutputPort())
mc.SetValue(0,2)
skinMapper = vtk.vtkPolyDataMapper();
skinMapper.SetInputConnection(mc.GetOutputPort())
skinMapper.ScalarVisibilityOff()
skin = vtk.vtkActor()
skin.SetMapper(skinMapper)
aRenderer.AddActor(skin)
iren.Initialize()
iren.Start()
The example that you have found is for visualizing isosurface of a grayscale volume. To display a labelmap volume in 3D, you need to use discrete marching cubes (or flying edges, for faster operation), set multiple values (SetValue), enable ScalarVisibility, set an appropriate color lookup table, and apply surface smoothing. See complete example here: https://lorensen.github.io/VTKExamples/site/Cxx/Medical/GenerateModelsFromLabels/
Thanks for your reply!!!
However, I don’t know how to display the image using Python. The classes in the example just tell users how to write the image into ‘.vtp’ files.
v16 = vtk.vtkNIFTIImageReader()
v16.SetDataByteOrderToLittleEndian()
v16.SetFileName(path)
mc = vtk.vtkDiscreteMarchingCubes()
mc.SetInputConnection(v16.GetOutputPort())
mc.SetValue(0,1)
smoother = vtk.vtkWindowedSincPolyDataFilter()
scalarsOff = vtk.vtkMaskFields()
selector = vtk.vtkThreshold()
geometry = vtk.vtkGeometryFilter()
smoother.SetInputConnection(mc.GetOutputPort())
smoother.SetNumberOfIterations(15)
smoother.BoundarySmoothingOff()
smoother.SetFeatureAngle(120)
smoother.Update()
selector.SetInputConnection(smoother.GetOutputPort())
selector.SetInputArrayToProcess(0,0,0,vtk.vtkDataObject.FIELD_ASSOCIATION_CELLS,
vtk.vtkDataSetAttributes.SCALARS)
scalarsOff.SetInputConnection(selector.GetOutputPort())
scalarsOff.CopyAttributeOff(vtk.vtkMaskFields.POINT_DATA, vtk.vtkDataSetAttributes.SCALARS)
scalarsOff.CopyAttributeOff(vtk.vtkMaskFields.CELL_DATA, vtk.vtkDataSetAttributes.SCALARS)
geometry.SetInputConnection(scalarsOff.GetOutputPort())
I’m using Python to solve the above problem but it doesn’t work…