Hi, everyone I am attempting to convert 3D numpy array into volume rendering function in pythonVTK or C++ code of VTK. Data type conversion of numpy array data into 3D volumes is an issue for me, I cant seem to find an alternative to file formats or conversion to data file formats of VTK itself.
Here is a full example of how to create a volume (vtkImageData) from a numpy array. The code supports scalar, vector, and tensor volumes. If you don’t use this from 3D Slicer then you need to change how you set the image spacing (you need to set it using vtkImageData.SetSpacing):
Note that you can get static or interactive (you can rotate around with your mouse) 3D volume rendering from numpy arrays directly in your Jupyter notebook by choosing 3D Slicer as kernel.
PyVista makes this easy:
import pyvista as pv
data = pv.wrap(my_3d_numpy_array)
data.plot(volume=True) # Volume render
See:
Can I display this on a webapp, somehow?
Yes, you can create a web app using Jupyter notebooks and Voila.
If you need interactive rendering (so that you can rotate the view with your mouse, etc.) then you can use 3D Slicer’s Python3-based Jupyter kernel, which already bundles VTK. It would not be too much work to set this up to work with PyVista either.
@banesullivan you could consider adding an interactive Jupyter widget for PyVista, similar to what we implemented for Slicer. It is a mini remote renderer based on ipycanvas and ipyevents. It is really simple (less than 300 lines of code, fully implemented in Python) but it is a game changer, as it it makes available full power of desktop VTK rendering interactively in web applications or notebooks.
What is volumeNode?
Volume node is a thin wrapper container class, which contains vtkImageData and image direction. This was necessary, as until very recently, VTK could not store image direction in vtkImageData. The class also allows associating image data with information about how to display and store it. You can find more information about MRML nodes here.
Thank you for the help and we had success making it work hopeefully we will make some improves overtime.
@lassoan, thanks for pointing me to that!! It was really easy to get a prototype up and running with PyVista. I put it here: https://github.com/Kitware/ipyvtk-simple - this is very much a prototype and much more is needed to make it available for PyVista users
Looks great!
To make sure more people actually give it a try and realize just how powerful this is, I would suggest adding a launch | binder badge. Clicking on it would open a Jupyter notebook preconfigured with VTK, pyvista, and this interactive widget.
Hello,
I also try to convert a 3D numpy array into VTK, but I do get some bugs. The array is an computed mask image from the original ultrasound image. The mask I want to apply over the original image in order to only show the region of interest in the ultrasound. I am able to visualize the original 3D volume, but applying the mask gives me a result with gaps.
For now I’m using a vtk.vtkImageImport() and np.transpose() for the array, with SetDataScalarTypeToUnsignedChar(), SetNumberOfScalarComponents(1), SetDataExtent(extend of original image), SetWholeExtent(extend of original image), SetDataSpacing(spacing of original image).
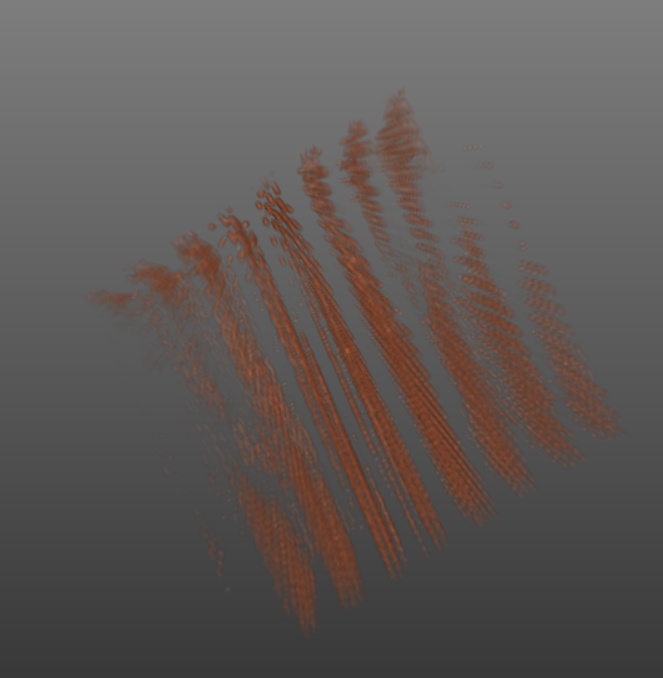
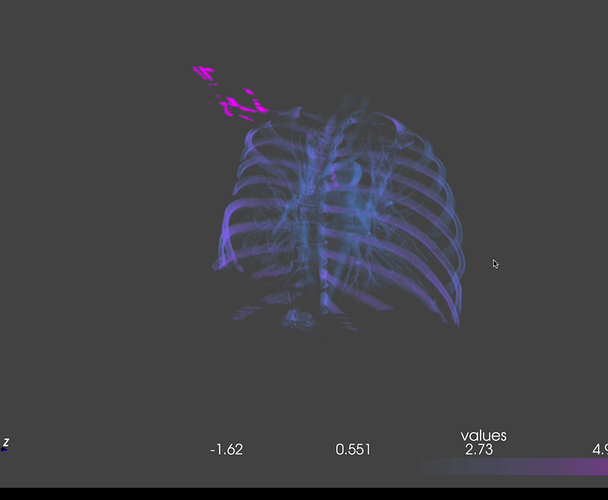
The output looks like this:
should I use vtkImageData? the gaps has to do something with the np.transpose() or how I set my dataspacing, but I tried a lot of things and this is the best result I got so far…
That looks like a classic F vs C ordering issue. Whenever you ravel/reshape the NumPy array and send it to VTK, you should play with the order (try F)
Otherwise, we’d need to see the actual code you are running
We made this function not sure if it helps
def displayer(numpy_mask):
data_matrix = numpy_mask
opacity = [0, 0, 0, 4, 8, 0, 0]
data = pv.wrap(data_matrix)
pv.set_plot_theme("night")
#print(type(data)) #pyvista.core.grid.UniformGrid'
#print(dir(data)) #x = pickle.dumps(data) #print(x)
#print(BytesIO(data))
return dataHi, could you please explain how to set the spacing? I can’t seem to figure out what volumeNode is and how to use it in my VTK pipeline.
Volume node is a thin wrapping over vtkImageData. Volume node stores an image and image origin, spacing, and axis directions (because VTK did not support image direction matrix until 1-2 years ago).
Standard numpy arrays don’t store image origin, spacing, and axis directions. If you need a numpy array that can store image metadata then you can have a look at xarray. Which is supported by a number of medical image computing packages, such as ITK.
@dgobbi have you looked at xarray and how ITK interfaces with it? It could be nice if the same Python image object could be used by ITK and VTK.
That’s great, thank you very much for your reply. I will look into ITK and xarray.